


Reproducible Manuscripts with Quarto - posit::conf(2023)
Presented by Mine Çetinkaya-Rundel In this talk, we present a new capability in Quarto that provides a straightforward and user-friendly approach to creating truly reproducible manuscripts that are publication-ready for submission to popular journals. This new feature, Quarto manuscripts, includes the ability to produce a bundled output containing a standardized journal format, source documents, source computations, referenced resources, and execution information into a single bundle that is ingested into journal review and production processes. We'll demo how Quarto manuscripts work and how you can incorporate them into your current manuscript development process as well as touch on pain points in your current workflow that Quarto manuscripts help alleviate. Presented at Posit Conference, between Sept 19-20 2023, Learn more at posit.co/conference. -------------------------- Talk Track: Quarto (1). Session Code: TALK-1070
image: thumbnail.jpg
Transcript#
This transcript was generated automatically and may contain errors.
Okay, thank you very much. Welcome to the Quarto session, and I have the distinct pleasure of talking about things that look like a stack of crepes, but actually they're manuscripts. So today I'm going to be talking about reproducible manuscripts with Quarto.
So I want to talk a little bit about the full sort of spectrum of complexity of reproducible scientific projects. What do I mean by that? Majority of my time as an educator particularly, and an educator who tends to teach students who are learning computing for the first time and does so with Quarto, I live in this wonderful land. It is a fantastic place to be. You have a single Quarto document. You are sort of like coding in one language because that's the language we're teaching the students, and we have full control over sort of like everything that they're doing, including the tools that they're using to write their code.
And the type of things that they're producing, while quite in-depth for them, are things that where the code can all live in a single file, and they don't mind running it over and over again with each edit. You want to change a header? Fine. Let's just run the whole compute again, and things go pretty fast and pretty neatly. And we tend to get HTML outputs as a result, and we can publish them on the web.
While that's like a majority of the time I'm living and breathing, the rest of the time I am thinking about, well, we collect data from these students about how they learn. At some point, we're going to want to publish these results. So how do we go from the simplest case to things that get sort of progressively more complex? So from simplest to let's go to simple. We can again have a document, a single Quarto document with just some R code in it, and with the help of the Quarto journal articles extension, I can output things to PDF or Word, not just output to any PDF or Word, but a document that looks exactly like the document in the journals I'm submitting my manuscripts to expect them to look like.
But again, the code in here is sort of, I'm going to say simple enough. That's not to say the analyses are very simplistic, but to say that I'm okay re-rendering this document every time I'm making edits to that, and this is a happy place to be too. But science is rarely that simple. Typically, you're working with multiple collaborators, each with their favorite computing language and code editor. And you're in multiple stages of a project, each with their own level of feasibility of what can be rerun with each edit and what needs to be cached. You don't want to be cleaning your data every time you want to change your headers for example.
So a more complex project might look like this. Maybe you have a single Quarto document, but a bunch of notebooks and stuff laying around, and maybe everyone's coding in R in your team, and you want to output to something like a PDF or Word for a journal submission. Or things can get even more exciting, where you have sort of the diversity of the tooling your collaboration team is using increases, where we have even more notebooks, where little bits of the project are being done. Maybe we have now multiple QMD files, some of them running R code in them, and some of them running Python code in them. But ultimately, we need to bring all of this together into a culminating manuscript that we want to submit to a journal.
The current state of reproducible scientific manuscripts
We can, turns out, leverage the notion of Quarto projects for writing these fully reproducible scientific manuscripts. So I'm going to take an aside for a second and talk about a little bit of a mind shift for some of you, and at least for myself as well. Whenever I use the word notebook, in my mind, I'm always thinking about like Jupyter notebooks because the word notebook is in there. But ultimately, a notebook is a document that contains both code and narrative. So going forward, when I say notebook, I could be talking about a Jupyter notebook, or I could be talking about a Quarto document, where you again see your code and narrative in the same place.
So let's keep that in mind. And think about what the current state of affairs is. Most computational science is born in notebooks, and it dies. That's sort of not so nice. Maybe ends in PDF or Word documents. And what happens in between is that we have peer review and publication workflows that don't tend to support these notebooks as research outputs. And these more complex scenarios that I talked about involve a lot of like manual finagling at the end. And not in a devious way, but just at the end of the project, you're just trying to get it over the threshold to bring the project to this journal submission stage.
And it's during this process that oftentimes reproducibility is lost. Or at least take some sort of second seat to formatting requirements. I will at least be honest and say that I have certainly done the thing where I had this beautiful notebook that I got to 90% of what the journal wanted as a Word file. And then I manually edited that Word file. And then I put a footnote saying the code can be found in the GitHub repository. Choose your own adventure.
And it's during this process that oftentimes reproducibility is lost.
And ultimately, the final submission that the journals want or archive once rarely captures all these computations. I mean, sure, they'll happily take your supplementary documents. But they rarely archive those documents with your manuscript. And sometimes they're not necessarily even reachable. So these can live in supplementary materials sometimes. And there is no ensuring that that GitHub repo I link to in my supplementary documentation is going to be there a year or two years from now. So if you've tried to reproduce others' work so that you could learn from them and iterate on it, you probably have had difficulty trying to sort of gather up all the pieces needed as well.
So there is, you know, this is the current state of affairs. But also within the current state of affairs, there's lots of great minds thinking about, well, how do we make this better? This is not a new problem. And there's been sort of like initiatives, various initiatives over at least my academic career of sort of trying to move reproducible computational science forward. And one of the sort of the current ongoing projects is this Notebooks Now project that's funded through this Sloan grant and a grant from the American Geophysical Union. And the idea is to try to sort of make these pieces work well.
A roadmap for fully reproducible manuscripts
So what could a roadmap look like for fully reproducible scientific manuscripts that are not just PDFs that are the outputs of a single QMD file? So what we need is one piece of the puzzle is we actually need an actual publishing workflow, an end-to-end scholarly publishing workflow, starting with that first notebook you started, you know, iterating on when you have the idea for your project to the final manuscript that you produced that treats things like Jupyter and Quarto notebooks as the primary element of scientific record, not just that PDF output.
We also need a publication process that sort of elevates transparent and reproducible workflows by authors where they are able to sort of bring together and submit and archive their software along with their document and their data. And this is going to take obviously lots of doing on the publisher's end, but it is putting on a bunch of new work, good work, good scientific work, but new work on the authors as well. So hopefully along with these initiatives come sort of new forms of credit to the folks who are working on this as well.
I'm going to focus on the first two pieces of the puzzle where Quarto can provide technical solutions to that and maybe perhaps then provide bandwidth for others to work on the third piece of the puzzle as well. So with Quarto, it can be authored in your favorite code editor so you no longer need to, you know, convince your collaborators to change their home. You can render QMD files or Jupyter notebooks to a variety of outputs. You can execute code in R, Python, and more. You can apply journal style to your outputs with these Quarto extensions and publish to, you know, a variety of venues including GitHub Pages, Netlify, and more.
Introducing Quarto manuscript projects
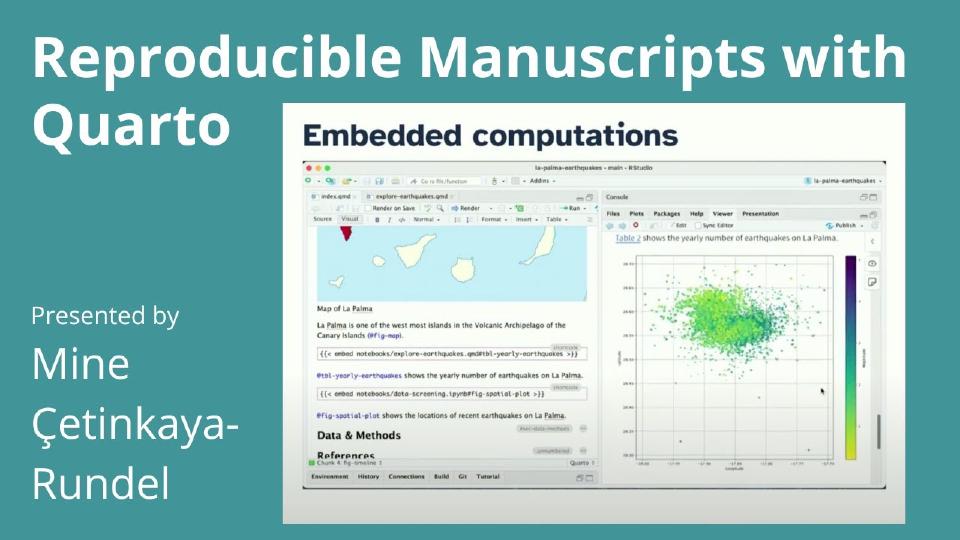
Now last year I said all of these things at this conference and over the year I hope that many of you have tried these things. And with Quarto projects we know that we can sort of orchestrate this notion of multiple inputs and multiple outputs. Well now with a new project type, that's the manuscript project type, you can orchestrate these multiple inputs and outputs again but you can also leverage this idea of embedded computations that I'll talk to you about in a little bit.
So let's introduce this new project type. This is Quarto 1.4 onward so there are pre-release versions available right now on the Quarto website but it is not available yet on the release version. We can produce manuscripts in multiple formats and give readers an easy way to sort of peruse them by creating a website that goes along with it and we can publish the computations so that readers can both sort of take a look at the computations and even go a step forward hopefully and start interacting with the computations as well.
So let's go ahead and write our manuscript. You can start any of these Quarto projects in one of two ways. You can say well create an empty project for me either using the RStudio IDE is usually what I tend to do or you can use the command line interface and do something like Quarto create project manuscript my favorite paper for example. Or go to the Quarto documentation and grab one of the projects that we've already sort of like prepared for you so that you can see all of the pieces working together and start replacing the bits of the narrative that don't relate to your work with your data and your narrative.
And if you do this in a folder that is being tracked by Git and has a GitHub repository associated with it you can have pretty easy going with your publishing as well. So with one command like Quarto publish you end up with a website that looks like this for your manuscript.
Demo: the manuscript website and features
So what we're seeing here is a variety of things that you would expect from a manuscript. We have figures that are actually cross-referenced and if I hover over these I can sort of see the cross-referenced figures as well. I can see that the articles are the notebooks where the computation is happening is sort of linked on the side and one of my favorite things that's sort of like on by default is you can go ahead and start doing things like highlighting the text.
And also if you have you can set up a group with your collaborators and say something like hey maybe we want to phrase these things slightly differently. So this hypothesis feature is turned on and so you no longer have to just like be doing those PDF annotations you can start interacting with things in this way.
Another thing is that from this one single source you get multiple formats. So on this rendered web page you can see that we're referencing to a PDF that has the AGU format so has a journal format as well as a Word file. So starting with the same YAML of my project right I have one YAML that says I want to HTML, I want a PDF, I also want a Word file. I can actually get to this output where I'm able to see and access each of these formats. In addition I get really rich front matter. So you're probably used to you know YAMLs that are a few lines long. This is like a whole novel in and of itself this YAML. But the need for this is that some journals want some things and other journals want other things and it's nice to be able to sort of put everything on there.
Each time you start a project you have a standard YAML that you fill with your collaborators and if you're submitting to this journal then only the relevant pieces will be picked up and rendered there and for the Word output that we were able to access from that home page again only the relevant pieces get picked up and fit in there as well.
Embedding computations from notebooks
For the next bit I'm going to sort of show you a video of how we can actually not sort of break this paradigm of a single Quarto file but also grab computations from notebooks as well. So I have a Quarto file and the data set that goes with it as well and let's take a look at the contents of that Quarto file. I'm housing all of that in the same folder as my manuscript and if I go ahead and render it you'll see that this particular notebook is not like my paper. It's just a few things I was trying, maybe one table or one figure I have created and let's assume for a second that there was a lot of compute required to create one of these.
Then I go to my main document, my manuscript, and I start using an embed shortcode to say I'm not going to bring in that code chunk here. What I want to do is I want that code chunk to stay in my notebook but I'm going to embed it in my main manuscript. So I'm able to grab the label of that code chunk and link it here.
And let's go ahead and take a look to see that the table that we produced in our notebook is embedded with a link that can take me back into the notebook as well. I had some R code in this document. My main manuscript was a QMD and I embedded a QMD here as well. This is also a nice opportunity to show you some of the nice features of the visual editor in RStudio. I'm able to create these cross references in a pretty straightforward manner and basically include them in my paper.
Now let's imagine that while I am a person who likes to write code in R, I have a collaborator who has created this fantastic visualization in their Jupyter notebook using Python code. So I've now received their notebook where they had already executed the code. And what I can do here is, again, I can now embed in the same QMD file output from that notebook as well. So I'm able to bring in outputs from different languages and different types of notebooks into the one manuscript file and not having to re-render that code again or re-execute that code again, but simply grab the results that were already created when my collaborator made that visualization for me.
And I can, again, using the same syntax, using the same syntax, do a cross reference to that as well. So let's go ahead and convince ourselves that that actually works. If I scroll down, I have my lovely visualization made with Python. I can go to the QMD file and start interacting with that notebook, or I can go to the IPython notebook and start interacting with that as well from this single document.
What's next: interactive computations
So what is next after this? Well, what I demonstrated here is that you can start perusing the code, which is fantastic. And at times, that's perhaps all you need. In fact, when I was first starting to play with the manuscripts feature, I was thinking, I wish it would just pop out and let me take a look at it and then go back to its home. I want to convince myself as to how that figure was created, but maybe I'm going to focus on reading this paper for the remainder of the time.
When it comes time to actually perhaps taking somebody else's paper and saying, I really want to understand how they implemented this method or how they created this table, I might want to genuinely dive into the code. And sure, I could go to the supplementary information and go to their GitHub repo and clone that repo and get started. But we know that there are a lot of hurdles along the way when you do that. So I want to interact with that code in a computational environment that's just a click away that has all the software and packages needed to reproduce the manuscript.
So I want to interact with that code in a computational environment that's just a click away that has all the software and packages needed to reproduce the manuscript.
In 2019, there was this article published in Nature where actually an article on eLife allowed you to do this. As of today, that article is paywalled, and you can't access the links anymore. And this is not to say the authors did anything wrong. It's to say it's been years, and this is very near and dear to a lot of scientists' hearts problems that's really difficult to solve.
So I think with the manuscript feature with Quarto, we're making advances into getting there. So as of today, you can start kicking the tires on the Quarto use binder feature, where you can see in addition to these static outputs, you have a link to be able to launch a binder instance. You do need to be a little bit patient as it does its thing there. But we would love for you to sort of go in and take a look to see how this is working for you. Is it capturing everything that you need? And is it working out for you? And give us some feedback on that as well.
So where do you go from here? I would recommend rewind back to that get started and start again. And I would point you to the documentation for the Quarto pre-release, where Charlotte has actually created a wonderful repository that you can get started with. Thank you so much for listening. And I'd be happy to take questions.
Q&A
I'm back. Thank you for the great presentation. So I'm going to turn things real, real quick with one question. When we submit a manuscript, we often don't do it just once. Do you have any tips for a workflow, like if you have to switch templates because you're submitting to a different journal? And or resources slash ways to contribute to the pool of journal templates that are already available?
Yeah. Wonderful. So it's actually that first bit is easier than you think. And I know that we don't tend to write an article for multiple journals at the same time. So it generally tends to be iterative, where you maybe submit to one and then you try another. But you can add other templates here as well. So in this case, I only had on line 10 here one of the journal templates being used. But you can add other ones here as well. So I could have added something like PLOS PDF or JASA PDF and kept going. And Quarto will happily render all of them for you, grab the necessary bits of the YAML, and create those PDFs for you.
There is a rich and growing sort of ecosystem of these extensions. And I think the best way to contribute, if you have a journal you want to submit to, is grab one of the ones. There's actually a starter repo in that Quarto journals templates organization as well. And start sort of like playing around with it. You may or may not, if you need a PDF output, you probably want to be sort of a little bit confident with writing a bit of like latex styling code. But I would also say that if you've started with a template for a journal of interest to you, but you're running into roadblocks, opening some issues and asking for help would be great as well. As the developers are often like watching that space and trying to help people get over that hurdle. Awesome. Thank you very much.




