

Hannah Frick - "Censored: A tidymodels package for survival models"
10 November 2022. "Censored: A tidymodels package for survival models" Hannah Frick, Software Engineer and Statistician, RStudio. CANSSI Ontario Statistical Software Conference. Abstract: Survival analysis is an important field in modeling, and there are many R packages available that implement various models, from "classic" parametric models to boosted trees. While they cover a great variety of model types, they also come with considerable amounts of heterogeneity in syntax. The tidymodels framework is a collection of R packages for modeling and machine learning using tidyverse principles. It provides a consistent interface to a variety of modeling functions, along with tools for resampling, assessing performance, and hyperparameter tuning. The censored package now extends the model coverage of tidymodels's parsnip package for survival analysis. It offers a tidymodels interface to parameteric survival models, (regularized) proportional hazards models and various tree-based models such as decision trees, boosted trees, and random forests for survival analysis. Additionally, it offers predictions of time to event, linear predictor, survival probability, or hazard in a tibble format consistent across different models
image: thumbnail.jpg
Transcript#
This transcript was generated automatically and may contain errors.
Thanks for having me. So yeah, I am working with censored regression survival analysis in the tidymodels framework. And when people hear survival analysis, they often think of like data from situations like this. And I mean, it is called survival analysis for a reason, but I'm going to include a friendly scheduled reminder that time to death is like not the only type of data where you might be interested in tools from sort of survival analysis.
My favorite little example has to do with newspapers, sort of a newspaper company is putting their papers out for people to grab in various locations. And they sort of want to know what the demand is there so that they can put the right number. And if at the end of the day, all of them are taken, they have a censored observation and it is not time that is censored, but the demand of the newspapers. So that helps me remember that sometimes you have censored data where you might not at first expect it. And I'm here because we are extending support for that type of analysis in tidymodels.
What is tidymodels and parsnip?
tidymodels is a framework and a collection of packages for modeling and machine learning, which uses tidyverse principles. And the part of tidymodels that's specifically for models rather than other elements of that whole modeling process is parsnip. And it is designed to give you some consistency in the interface. So consistency in how you specify and fit various types of models and in how you predict with them and also what you get back. So not like a hodgepodge of vector, matrix, array, data frame, that kind of thing. So that's what parsnip does. And censored, which was written very big on my title, is a parsnip extension package specifically for survival models.
So I'll talk largely about those two bullet points, how design principles for parsnip play out for censored regression and survival models in the censored package, how you specify and fit, and how you predict. And then I'll give a bit of an outlook beyond the models. But let's start with that first one.
Specifying and fitting models
So in parsnip, you need three components to do a full model specification. One is obviously the model type. So here is the first line, ran forest for a random forest model, could be a linear regression model, could be other things. Apart from the model type, you need to tell us how you want to fit that, which kind of implementation you want to use. In this case, it's the ranger R package. It does not have to be an R package. Most of the times it is. But sometimes other implementations can be used, like stand for Bayesian models, for example. And the third element here is the mode. So a random forest could be used for classification, for regression. So in order to have a full specification, you need to be clear about that and set the mode.
And these three elements are like a cornerstone of parsnip. So of course, they show up in the extension package censored, which packs the questions all right. What's actually new and different? We had to make a few additions to this, all for survival analysis. So for one, there's a new model type, just the proportional hazards model. Most famous one of those, the Cox model. There's a new mode to be explicit about you working with censored data. So that's the censored regression mode. And yes, you guessed it, the third element that's new are a bunch of new engines, partly for existing models, but always for that new mode.
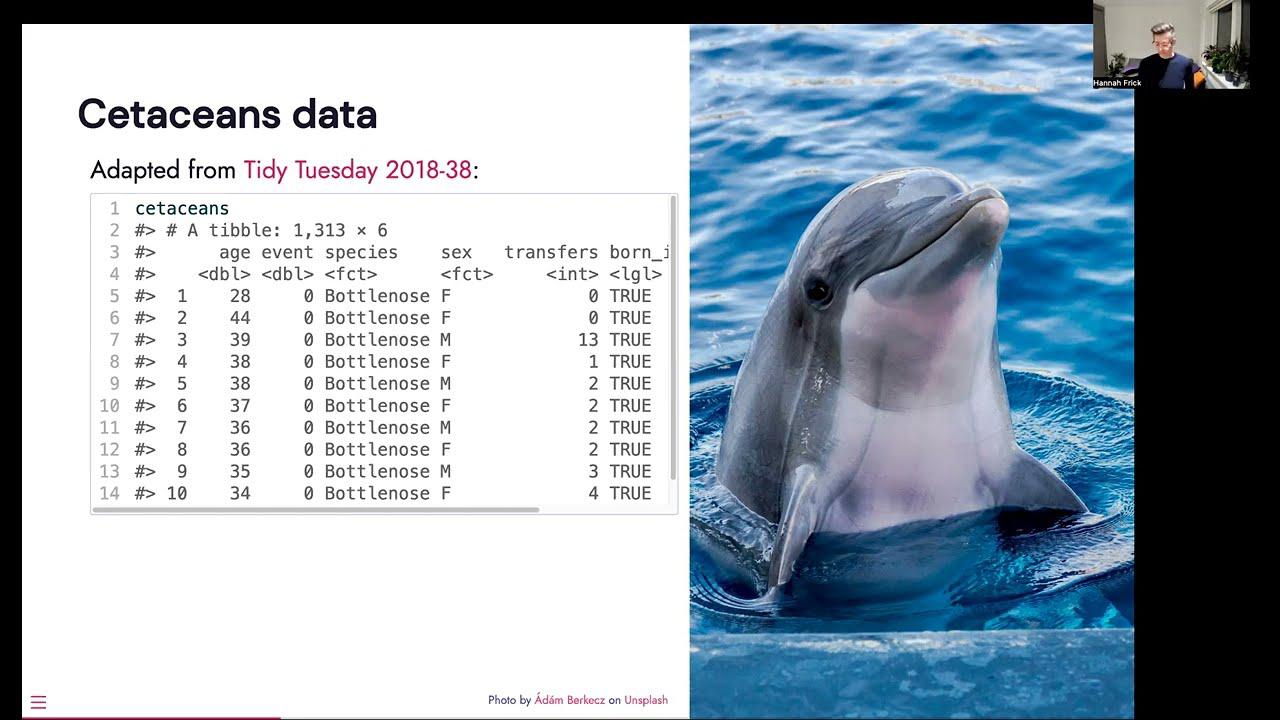
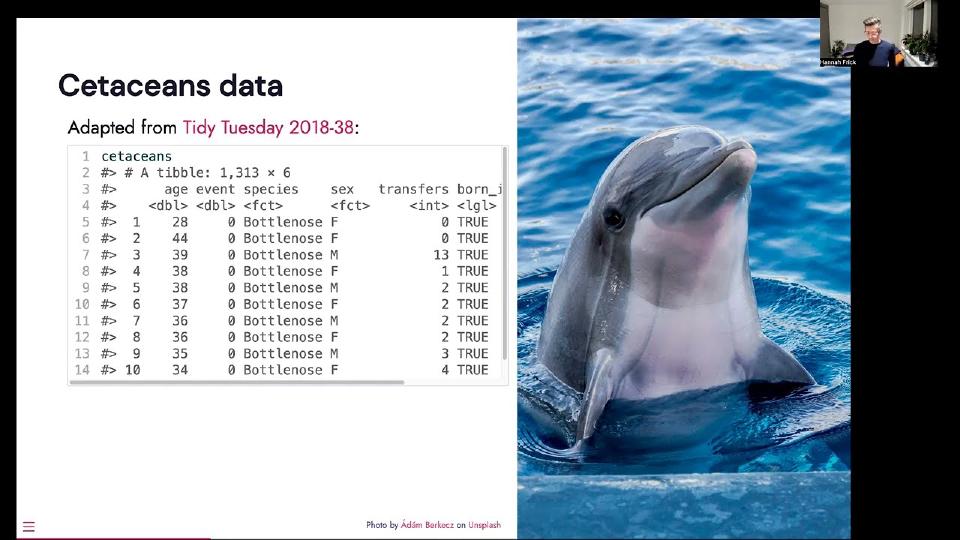
So they cover parametric models, semi-parametric models like that proportional hazards model and various tree-based models. And what is also in censored is the formula interface to all of these models, but it includes a way to specify stratification. And I bring some data, not New York subway, not newspapers, but I can offer some dolphins and whales living in captivity in the U.S. It's adapted from a tidy Tuesday data set. And we have a column for the age of the citation, an event indicator, whether they're dead or alive, species, sex, the number of transfers between different facilities in the U.S. and whether or not they were born in captivity.
So for doing this, it helps to load the censored package. And then we're going to use that new model type of proportional hazards model, use that new model type of proportional hazards model, and I'm going to set the engine to the survival package, fit a classic Cox model here, and set the mode to censored regression. That gives you the specification of what you generally want to do. That is not the fitted model yet. That's what you do with fit, and you give it your model formula, and you give it the data, and then you run it. And if you want to include stratification, you can do that as you're used to in the survival package and add a strata term in your model formula.
So that altogether gives you how to do that in censored. And if you're used to working with the survival package directly, you look at this and think, oh, that's a lot of lines of code. Yeah, it is. So if you're doing this just like survival package directly, it is more compact. You just got your CoxPH function, the formula, and the data, and you're good to go. And if that's what you want to do, that's good, and then I would probably also do this. So censored gets interesting if you want to try out that model, and then you might want to do something slightly different or completely different.
And to demonstrate that, I'm going to change from the classic Cox model to a penalized version of this. So I've just moved the code to the left, same as before, to make some room on the right to do this with the glimnet package. And the limit does not have a formula interface. It wants matrices. So we're going to make these our particular matrix x, and the response, which is that serve object. And if we want to do stratification, at glimnet, you have to stratify your response. So that's what we're doing next. And then you can take these two glimnet functions, set your family to Cox, set your penalty parameter to lambda, and then you're good to go.
And of course, this isn't the end of the world, and it's not that much to do, especially in this example. But if you're doing that a lot, I would argue that it gets a little bit tedious, especially sort of the part of having to change in what way you store and provide your data. So this is the kind of thing we wanted to make easier. So it looks a lot more boring if we do this with censored. I've taken what we've done already on the left, copied it over to the right, and then the main thing I've changed to go from the regular Cox model to the penalized version is to change the engine. I want to fit the same model basically one time with the survival package and one time with the glimnet package. The other thing we need to do is tell the specification what the penalty parameter is. The other two parts, like the mode and how to fit that, and what is the response and what are the predictors, that actually is the same.
So this is the kind of thing we wanted to make easier. So it looks a lot more boring if we do this with censored.
So the main part is sort of the model type and the engine. And if you want to go to something rather different than this, say a boosted tree, then you have to change essentially two lines. So now the model type is a boosted tree, and we're going to use the mboost package as the engine here, and the rest is basically the same. So that is the idea to give you a certain amount of ease moving from one to another if you need to try out more than one.
So what is all there? That's a little table of the model types and the engines that we have. They're all for that new mode, sensor regression, at the bottom is like survival reg for parametric models, proportional hazards you've just seen with the two engines, and then the tree-based methods that I mentioned are decision trees, random forests, back trees, and boosted trees.
And new in the development version, we have another engine, FlexServeSpline, for making parametric survival models with a spline-based distribution. This has been contributed by Matt Parkinton, so many things for that. And the other new engine has also been a contribution by Byron Viego. That is for the model type random forests, but the engine is one that calculates accelerated bleak random survival forests. And if you want to try that out, you can install the development version from GitHub.
Predicting with survival models
And that's sort of the part on how do specified fit models are censored. So let's get to the predict part. And also here, there's an element of what is already in tidymodels and how that then plays out here. So predictions through tidymodels code will adhere to these three principles. For one, they'll always be in a table, so you know what kind of data format you get. Table as the type, but you will also know the column names and types of this. They will be unsurprising. And the third element is that the number of rows in new data, so the data that you predict on, and the output of your call to prediction are the same. Sounds pretty unassuming. That's relevant. If you have missing data, and you will get an NA value for those observations rather than them being skipped and removed, and then you might have to figure out where which observation goes if you want to merge it back to your original data or your true values.
So that's stuff that comes from the existing tidymodels framework. So what is new for survival analysis? The answer is new prediction types. So what you can predict with these models is, for one, survival time. The expected survival time. I'm just taking the first three rows of the citations data set. I'm going to add an NA in the first row to make things a little more interesting. And then we call predict. We give it the model, we give it a new data, and we set the type to time. We get back. The column that you get for this type will always be named dot red underscore time. And you have three rows because you put three rows in.
The other major prediction type here is survival probability. That's the type. Survival here, line four. But survival probability is always a probability at a certain point in time. So we are calculating that for age 10, 15, 20, and 40 years for these citations. And we get back a table with three rows. This is called dot pred. But we ask for four time points. So these are stashed into little tables inside that where you have the time points. So if we pull the first one out, you see the column dot time with the four time points that I asked for. And dot pred underscore survival is all in A because I snuck in an NA. If that's not the case, then you do get predicted probabilities.
What sensor does not give you is a surfit object in order to describe survival curves. If you want to say plot that for illustration purposes, you can just amp up the number of time points to cover your range. So I'm going to go for one year to 80 years. And then you can unnest the thing that will stack all the tables, the inside tables underneath each other and make one big table. And because I want to keep track of who's who, I'm sneaking in the mutate statement here. But that sort of long table can then be passed on to ggplot and you put a GM step over it to give you a good illustration of how the survival probability unfortunately goes down over time.
So the prediction types in sensor that you can use are the survival time and the survival probability. You can do that for all of the models that we provide there. And then depending on the engine and what the engine provides, you can also have predictions of the hazard, quantiles of the event time distribution or the linear predictor if your model has one. And for all of these, you don't have to do your own prep for a matrix interface or pad results for a maze or sort of any of the other things that you might have to.
Beyond the models: workflows and the tidymodels framework
And that's the main point for prediction. And now I want to take you on a little tour of sort of peeking beyond the models. I mentioned that tidymodels is a framework covering the whole process. So there is functionality for preprocessing for the models, which we've talked about at length here. Workflows to combine those two into one object. Resampling, performance metrics, tuning a la jazz. And I want to tell you that the first four are all fine to work with with survival models. And want to show you some of that in action, pointing out a few practical tips as we go along.
Maybe a little bit of background is like, what is that workflow thing you're talking about? And we would like to get you to think about the model auto model workflow, not just as the actual maybe a linear model or proportional hazards model, but to think of it as all the parts that need estimation from the data. So if you later want to assess performance of this and cross validate it, you need to do both of these steps. Everything where you do estimation on all of your resamples. So tidymodels provides a workflow object that bundles those two together and makes it easier to hand that off to the different parts. That is to keep you from having accidental information or data leakage because you estimate your principal components on all of the data, including stuff that is not used when you fit your model.
So the preprocessing, there isn't really anything specifically new that needs doing for survival analysis, but it is good to remember a general recommendation, which is to handle transformations of the response outside of a recipe. So before you start making your recipe, you do that one step. So for survival analysis, that means make that serve object, which is typically a response, make that at the start. So I'm taking the citations data set and put it through a mutate, make the serve object and discard the age and event column because obviously the information is in the serve column now.
So after doing this, we're going to make a workflow with a model and the recipe. The model is still the proportional hazards model. I'm going to go with the Glennet, the penalized version of this. If you do that, it works better if your predictors are normalized. So we're going to make a recipe during that, making some dummy variables for Glennet, normalizing all the numeric predictors, and now we're going to put those two in a workflow. That's just a call to workflow. You add a recipe and then you add a model.
And then the only like, oh, moment that you might have in this is that if you're used to working, um, like the base formula as a way to, uh, do kind of preprocessing, like turning factors into dummy variables and all that stuff, um, and specifying very specific things about that model, like a stratification, um, then it's good to remember that preprocessing recipes is defined by the steps and the formula that you put in at the very start is only used to assign roles and get the data type so that you can do selectors like all numeric predictors. Model specific terms like strata can only be understood by the model, not any model, but like the survival model. Um, so that's why you need to keep it with the model. That's why it is in a, in the call to add model as a formula argument. That's where you can specify that. Um, it helps me to think of like this as a model formula as opposed to a preprocessing formula.
But if you stack these two things together, you can then work with that workflow object like you did with the model before. You can say fit on it and give it the data, and then you can take that fitted object and predict with it. So that's sort of working as usual.
What's next
I'm currently working on simplifying that workflows bit a little bit. The model specific terms will always need to be specified with the model. But if you have a formula that does not have that, like say remove that call to strata, then that should be a little easier. Um, and that's what I'm currently working on. And what we as a team work on are performance metrics. So we currently don't have survival analysis specific metrics. Um, there is a next like time-dependent RST curves and prior scores. Um, and that will then enable us to do a tight integration with resampling and specifically tuning. That's obviously without performance metrics, no tuning.
But to sort of tie it all together and go back to what the title was, that was the sensor package. And I'd like you to remember that it's a consistent interface to survival models and sort of gives you consistency in how you specify and fit and consistency in how you predict and what you get back. So please have a try, give us feedback. Thank you so much.









