What's New in Dplyr (0.7.0) | RStudio Webinar - 2017
This is a recording of an RStudio webinar. You can subscribe to receive invitations to future webinars at https://www.rstudio.com/resources/webinars/ . We try to host a couple each month with the goal of furthering the R community's understanding of R and RStudio's capabilities. We are always interested in receiving feedback, so please don't hesitate to comment or reach out with a personal message
image: thumbnail.jpg
Transcript#
This transcript was generated automatically and may contain errors.
Well hi everyone and thanks for joining me on this webinar. I'm going to do kind of a quick overview of some of the new stuff in dplyr 0.7. We'll start with kind of a bunch of smaller stuff, none of it's kind of particularly exciting by itself, but in aggregate it adds up to just relieving a whole bunch of small pain points in dplyr. Then I'll talk a little bit about databases and then finish off by talking about TidyEval, which is a new system for programming in dplyr. So there should be plenty of time for questions. I'll stop after each section and take any that look particularly interesting to the whole crowd.
Okay, so the first thing I wanted to mention, if you don't already know about it, dplyr has a website now, dplyr.tidyverse.org. That's a great way if you want to see the examples and the code, see the vignettes, and generally get an overview of the package. So that's a great place to start if there's anything you want to know more about dplyr after this webinar. I also wanted to mention briefly the Tidyverse package. If you haven't heard of the Tidyverse, the Tidyverse package just basically makes it easy to get a bunch of packages that work well together. So when you install it, it installs all of the packages in the Tidyverse, and then when you load it, it loads the most important packages.
So here I'm showing you the development version, which in conjunction with the development version of RStudio gives you this pretty colorful display telling you which packages are loaded. So you can see here I'm using the development version of Purr. It gives you a little bit of information, a little bit of session info, so you can, if you're having problems, you can, you know, if your code works on your computer and not someone else's, you can often track down the difference to one of these issues. And then finally tells you what are the functions that the Tidyverse is masking from base R or other packages.
New datasets
So with that, let's dive into some of the new stuff in dplyr. The first one is that there's two new datasets that, if you're teaching dplyr, might be handy, and we'll be using these more and more in the examples to illustrate more sophisticated features of dplyr. So the first one is the Star Wars dataset, which contains a bunch of information about the characters from Star Wars, which come from the SWAPI, the Star Wars API site. And one of the things that's different about this dataset compared to other datasets and other packages is this contains a few simple list columns. So for example, you can look at this films column, and that tells you it's a list column because each person may have appeared in more than one.
So Star Wars, some fun data about the characters in the Star Wars universe, and contains some list columns to practice your list column skills on. The other new dataset is this dataset about storms in the Atlantic. This contains a whole bunch of data from the NOAA Atlantic hurricane database. So there are 198 storms in here, going from 1975 to today. We can do a quick plot, and you can see approximately where they start out, and where they end. So this dataset has a rich grouping structure, because you're probably going to want to group each storm, and then compute various summary statistics on.
The pull verb
So two new datasets, there's a handful of other datasets that you can read about in the release notes, or the blog posts, that are just really simple datasets, if you want to teach people basic join functions, or practice them yourself. There's one new important verb in dplyr called pull. Pull gives you a single column. So this has been a long-time source of frustration for many dplyr users. If you use select, and give it a single variable name, it's still going to give you a tibble back. It's going to give you a tibble that just has a single column. If you use pull, it is going to give you that single vector.
This is basically equivalent to this code you could do already, using the dollar sign, but it works on database backends. And the other thing that's sometimes handy, is you can use it to select columns by their position. You can select from the front, from the left-hand side, if you give it a positive number, or you can select from the right-hand side, if you give a negative number. And indeed if you don't give pull anything, that is the default. It's going to take the right most column by default, kind of under the assumption that, if you've just created something, you probably want to take a look at it.
Error messages, encoding, and case_when
So that's pull. There's a bunch of, we put a bunch of work into improving the error messages, so hopefully if you do something by accident, that you didn't mean to, you should now get a better error message from dplyr, that tells you both where the problem is, i.e. the column, what it was expecting, and what you gave it, which would hopefully make any problems easy to diagnose in dplyr.
We also put a bunch of time into encoding support in dplyr. Probably the most, the case that affected the most number of people, was if you use CJK, Chinese, Japanese, or Korean languages, or data sets containing those languages on Windows, you basically could not use dplyr without a huge amount of pain. So that should now all be resolved, and if you discover any differently, please let us know so we can fix that. We also made some changes to make joins and similar functions behave better when the columns you're joining have different encodings.
I just wanted to still illustrate one little example of this, just to illustrate how painful encodings in R can be. Here I've just created a string, that's the word I live, which if I remember my high school French correctly, means student. And when you print these two strings out, they look identical, and indeed if you ask identical, are they the same, it says yes. But if you go and say, well tell me the binary representation of the string using char to raw, we can see that they're actually different strings, and that's because they have a different encoding. So you can see X is encoded as UTF-8, and Y is encoded as Latin 1.
So now dplyr should just kind of handle this problem for you automatically. If you're using Redar and other packages in the tidyverse, you should always get UTF data as your input, generally that's easier to work with, just stick with UTF-8 rather than messing around with different encodings. So whatever your encoding is on disk, load that up, convert it to UTF-8, and use that in dplyr.
One other tiny little feature that's quite nice is now that case when function works inside mutate. Maybe you have not seen case when before, but it's a really useful alternative to a whole bunch of nested if-elses. So I'll show you a quick little demo of using case when, just a simple case, and then show what that looks like now that it works inside mutate.
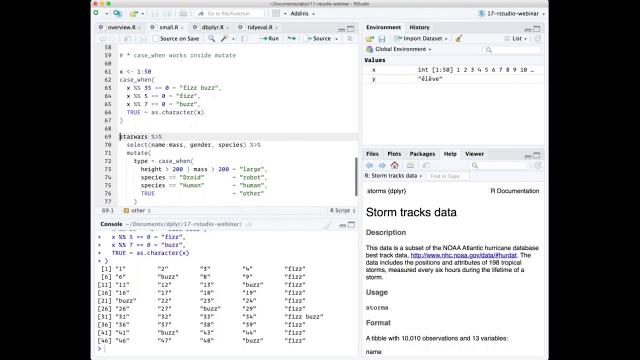
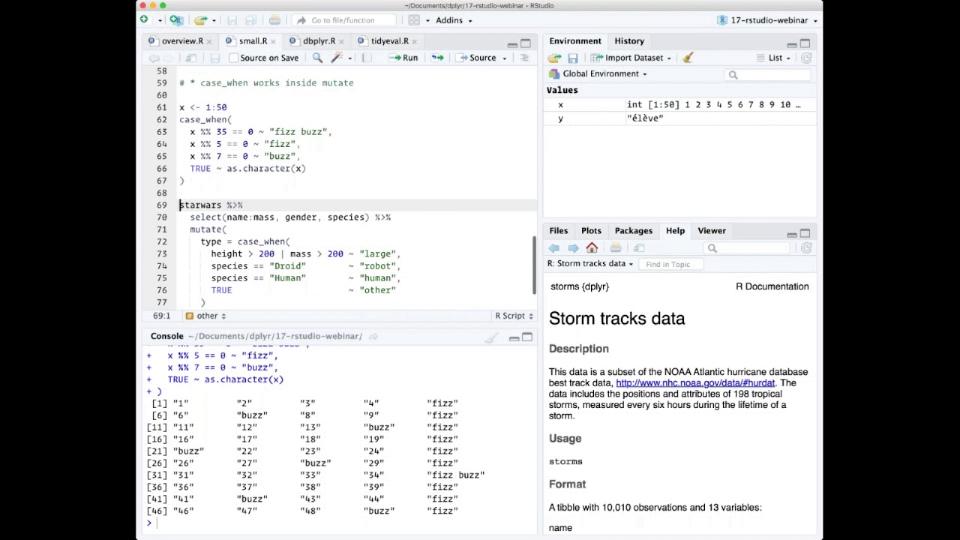
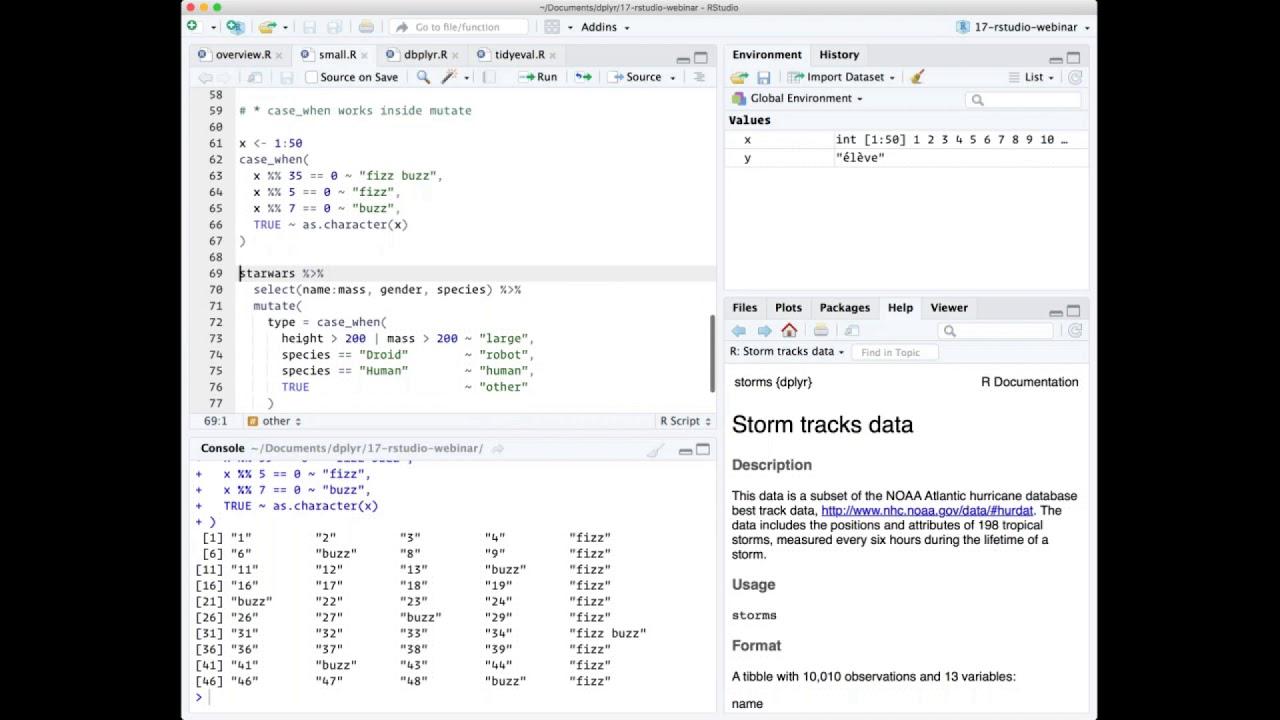
So here I'm going to use case when to implement the sort of infamous fizzbuzz programming challenge. And basically in the fizzbuzz programming challenge, what you have to do is print out a vector of 50 or 100 numbers, but when that number is divisible by 5, you need to print fizz. When it's divisible by 7, you need to print buzz. And when it's divisible by 5 and 7, you need to print fizzbuzz. In case when allows you to express this very succinctly, it says basically we use the formula, the tilde is basically kind of an if-then. So if it's divisible by 35, then return fizzbuzz. If it's divisible by 5, then return fizz. If it's divisible by 7, then return buzz. Otherwise, for everything else, true is always going to be true, convert it to a character. So case when operates sequentially, so as soon as the first thing is true, it's going to ignore the rest.
A more realistic use is to use it in the context of a data frame. It's really useful when you need to recode variables. So here I've just kind of made a very somewhat silly example using the Star Wars data set where I want to categorize the different characters based on a combination of their height and their mass and their species. So I'm saying if the height is greater than 200 centimeters or their mass is greater than 200 kilograms, I'm going to say that they're a large type. If they are a droid, I'm going to say they're a robot type. If they're a human, I'm going to say they're a human type. Otherwise, so true means otherwise, I'll just mark them as other. So when I run this, I get a new column called type which succinctly expresses exactly how I want to do that transformation. And this is much easier than doing the kind of equivalent set of if-else statements.
Database support
Okay, so next I wanted to talk about the new database support in dplyr or the improved database support in dplyr. Again, if you want to learn more about this, the best place is the website dbplyr. So dbplyr is the package that now provides all the database related functionality in dplyr. By and large, you will not need to know about this package. You will generally not need to attach this package, but dplyr will load that for you. You'll need to install it and you'll need to know that this is where that code is if you need to get help.
So the biggest change in this version of dbplyr is that there's much much better integration with the DBI package now. So the DBI package is a uniform interface to many, many, many different databases. So to use the DBI package, you always pair it with a database specific R package. So for example, I am going to create a connection to a database. So to create a connection to a database, you use the dbconnect function from the DBI package. And the first argument to that function is the database driver or the R package that provides the database connection. So here I'm going to connect to a SQLite database, which is provided by the SQLite package.
If you haven't heard of SQLite before, this is a fantastic place to start with databases because the database is actually included in the SQLite package. So most database systems, you've got like a database server, which can be a pain to set up and get running. With SQLite, that database is actually bundled in the package for you. So it's really, really easy to get up and going. And I'm going to cheat even further, I'm going to make this even easier to get up and going with, and I'm going to use a temporary in-memory database.
So I have a connection to this database, and you can use DBI to connect to all sorts of packages, SQLite, MySQL, Postgres. And then I'll talk a little bit later about some of the new enterprise databases that we're working to build support for at RStudio as well. So I've got a connection to that database, and now I am going to use one of the DBI functions, DBI write table, to copy a data frame to that database. And so what's new in this version is basically the seamless interaction between DBI, which is a very low level function that allows you to execute SQL queries directly, if you know SQL, and dplyr, which is a much higher level interface, which will write those SQL queries for you.
So now I can say, create a new table from this connection. So I print this out, you'll see that this is now a remote table. It's a table in a database, and in fact in that database is a SQLite database. And the neat thing about using dplyr with a database is that dplyr will generate the SQL for you. And there's been a bunch of improvements to the SQL generation in this version. One of the really nice improvements is this optimizer that was contributed by H. Hifflin, which reduces a lot of the sub queries that dplyr added previously.
So let's take a little example. I'm going to take this MT cars 2 data set, I'm going to filter it, I'm going to select some variables, and then I'm only going to look at the top 10. And if I just run this code, what dplyr does behind the scenes is generate the SQL query, sends that to the database, the database does all the work, and then returns the results. If you want to see the query that dplyr is generating, you can use the show query function. And you can see here this query is probably pretty similar to what you'd write by hand. We're going to select these variables from this table, where this condition is true, and I'm going to limit the results to at most two. Now in the previous version of dplyr, this would have generated at least, you know, three sub queries, which in some databases can degrade performance, and definitely makes it harder to understand what dplyr is doing when it generates the SQL for you.
We've also added more support for type coercion, so if you need to change the type of a variable in the database to a character, or a vector, or an integer, you can use the R functions you're familiar with, and those will automatically be converted behind the scenes to the appropriate SQL. So here you can see I'm using transmute. If you haven't heard of transmute before, it's a variant of mutate that just only keeps the variables you specify, it doesn't keep all the other variables.
Another thing that's changed in the latest version of dplyr is that there's an annoying bug whenever you're using in. Now this always generates the correct SQL. Previously we'd forget to put the parentheses in there, which is valid R code, but not valid SQL code.
So baked in to dbplyr, there's translation support for the major open source databases, SQLite, MySQL, and PostgreSQL. We are working to add more support for commercial databases. So this version also includes translation support for MS, Microsoft SQL Server, Apache Impala and Apache Hive when you use them through the ODBC package. The ODBC package is another package we've been working on recently. It is a DBI compliant back end for ODBC databases. So this makes, ODBC is a very popular standard for connecting to databases. This makes it very, very easy to connect to a whole bunch of commercial databases that are not, that do not have the best support in R right now.
So this is something we're working on hard at RStudio at the moment. You'll hear more from us in the near future, but we want connecting to any database. We want to make it as easy as possible to get your data from wherever it is, whether it's SQL Server, whether it's Oracle, whether it's Postgres, whether it's Redshift, to get that data from wherever it lives inside your organization and into R. And we've been working on a whole set of, there's like 10 packages involved in various ways at the moment. We'll be sharing that vision with you in the very near future.
But the other big thing that a lot of people have been clamoring for is support for schemas. So I'm just going to do a little demonstration of that. I'm going to create another database, another SQLite database, and I'm going to attach it. This is the way that SQLite allows you to work with multiple schemas. SQLite is kind of a very small database. It doesn't have like amazing schema support, but at least allows you to demonstrate the feature. I'm going to create two tables, both called df. One is going to be the iris data frame, and one is going to be the empty cars data frame. And I'm going to put these in different schemes.
So now dbplyr has this new function called in schema, which allows you to be specific about what schema you want to look for the table in. So here this is saying, look in the auxiliary schema for a table named df. And when we do that, what you can see here basically is that when we refer to this, we can now be explicit about what schema we want to get our table from. So if you don't use schemas, this does not affect you at all, but many big corporate databases use schemas extensively, and this should make your life much, much easier.
So if you don't use schemas, this does not affect you at all, but many big corporate databases use schemas extensively, and this should make your life much, much easier.
TidyEval: programming with dplyr
Okay, so let's finish off by talking about tidy eval, which is kind of the biggest and most complicated new feature in dplyr. And before I sort of talk about exactly what this is, I want to like illustrate the problem that we're trying to solve. And so here I've got a little table and I'm doing some data analysis with it. And basically each of these three clumps of code, I am doing the same summary, but with a different group. And so to do this with dplyr, you might be copying and pasting your code. And that's great. I think a rule of thumb is it's fine to kind of copy and paste up to like three times. But as soon as you go beyond that, it's worth the investment to write a function.
And that's because often, you know, the thing that you're doing changes, or you discover a bug. And if you've copied and pasted that code everywhere, tracking down every single instance of that bug can be really painful. So it's a really good idea to turn repeated code that you've created by copying and pasting into a function. But this is hard because dplyr lacks a property called referential transparency.
So let me try and just describe this quickly. I've created two simple functions, one that just multiplies its input by 10, and one that adds 10. And normally, when you're writing functions in R, you can extract out kind of a subextract expression or part of that function and assign it to a variable. And that doesn't matter. You can do that as much as you like, and you always get the same result. That's what's called referential transparency. You can create new variables, and it doesn't change how the expression is computed. But this doesn't work with dplyr because you can't pull out that expression. You can't pull out this repeated code, this G1, this G2, and just G3. You can't assign that to a variable because there is no objects called that. Because inside the dplyr expressions, to make your data analysis, your data manipulation as fast and fluid as possible, this variable refers to a variable inside the data.
So this is one of the features of R called the sometimes called nonstandard evaluation, where we're taking this expression and we're evaluating it in a special way. And this is really great because it allows you to save so much typing. You're not constantly having to type the name of the data frame again and again and again in your expressions. But the downside of this implicitness is that it's hard to program. And so in this version of dplyr, we have a new system called tidy eval, which is basically a new framework for programming with dplyr, dealing with this type of problem in general.
And so if you encounter this problem, you might say, well, I can't assign it to, I can't assign the bear variable name. Maybe I could try putting that inside a string. And that still doesn't work because it's looking for a column called group var. It's not looking inside that variable to find the G2. So to solve this problem, we have a new data structure called a quosure, which you can create with the quo function. So this basically kind of captures the expression. So this is what dplyr is all about. It's about capturing what you want and evaluating it at a different time.
So when you do this in dplyr, it's about capturing this expression and evaluating in a different context, in a context where you don't have to say which data frame every variable comes with. Or in the case of when you're talking to a database, the expression actually doesn't even get evaluated by R. It gets evaluated by the database much, much later on. So now we can use this quo function to kind of capture this variable. So we're saying we don't want, we want to use this variable G1 later on.
Now, unfortunately, that still doesn't work with group var. And what we need is some way to kind of unquote this variable. We just say, don't take this valuable, don't take this input literally, look inside this variable and see what it looks like. And so we now have this unquoting tool, bang, bang. And so what this tells dplyr to do is don't just say, don't look at group var literally, look at what's inside a group var and evaluate that. And so this gives us the ability to program with dplyr. We can now create functions where you can write functions that work like dplyr does. We can even take this further. You can not only do the grouping variables, but you can use this in every single function in dplyr, whether it's summarize or mutate or mean or whatever.
So this, I am like a hundred percent confident that the theory behind this is correct and robust and works like a hundred percent of the time, not just 95% of the time. We are still working to like try and explain it in a way that you can understand.
So this, I am like a hundred percent confident that the theory behind this is correct and robust and works like a hundred percent of the time, not just 95% of the time. We are still working to like try and explain it in a way that you can understand. This is basically, it's sort of a similar level of complexity to learning how to write functions. So if my explanation right now just left you reeling and confused, do not blame yourself. That is my fault that I do not know a good way of explaining that yet, but we are working on that. You can always see kind of our latest efforts on the dplyr website. There was an article about programming with dplyr that explains the problem in much more depth. And we will keep rewriting that until we are confident that we have figured out a good way to explain what's going on here in a way that you can understand and then deploy in your own code.
Q&A
So that's all I had to talk about. Again, I'm just going to kind of skim through and if anyone else from RStudio wants to like raise any questions to me, please do so. But I'll just take a quick look at these questions and see what might be of interest broadly.
So one question is, what's the difference between this new system and the previous system which used underscores after function names? It used functions and it used formulas. And basically the problem with the old system is that there was no easy way to write code like this. You'd have to do something. And I honestly, I don't even, I think you have to write like interp and then you have to do something like this, I think. So basically the previous system of non-standard evaluation was extremely constrained and it didn't actually help you solve the problems that you really needed to solve. So again, this new system, it's a little bit more complicated. It's going to take a little bit more work to understand what's going on. But I think that investing the time in that is really going to pay off because it allows you to do more things in dplyr.
And we're going to be rolling out the system everywhere in the tidyverse. So hopefully in the next month or so, you'll be able to use the same system in tidyr. And by the end of the year, you'll be able to use the same system in ggplot2. So again, it's an investment. It's going to take a little bit of time to get your head around it. But once you do, you're going to be using it in many, many, many places. It's going to make programming with all of these kind of slightly magical functions designed for data analysis much, much easier.
Yeah. So someone pointed out that quo and bang bang are kind of somewhat equivalent to B quote and eval. Yes, that is absolutely correct. Although B quote only handles the expression, not the associated environment, which leads to subtle bugs that again, sort of B quote kind of works like, you know, 95% of the time, which makes the 5% of the time it doesn't work really, really frustrating to debug. We've tried to move that pain kind of upfront with tidy eval. It's going to be a little bit more painful to learn it, but once you've learned it, there should be far fewer unpleasant surprises down the end.
Bunch of questions are about quo and in quo. Yes. So a lot of you spotted that I used quo in my example up here, but then I used in quo inside the function. The difference here is a little bit subtle, but quo basically always gives you its input back. So if I used quo inside this function, it's always going to put group var, but here I don't actually want group var. I want the value in the group var variable. I want to kind of go one level up to see what's inside that expression. And that's what in quo does rather than quo. So in quo captures exactly what's inside the parentheses. In quo you use inside a function that looks at that argument and see what expression was associated with that expression.
A couple of people asked about whether case when translates to SQL queries. That is a very good question that I will have to look at the source code to answer. It was definitely inspired by it. So it looks like we do not currently have a translation for it, but case when was inspired by the case when function in SQL. So I'm just going to quickly file an issue so I don't forget to add a SQL translation for case when. Translation for case when. So we'll make sure that happens in the next version.
So we had lots of questions about unquoting. Basically, there are, I kind of don't want to answer all of them here because they're fairly specific. But if you have any questions, do please read this programming article, which should hopefully answer your questions. If it does not, please let me know. You can shoot me an email or send an email to the manipulator programming list and we'll work. But basically, looking through the questions, if you did ask a question about this, the answer is yes, it is possible. It is similar to, it's kind of like macro variables in SAS, although those are just kind of work on the basis of text substitution. This is much more robust and works in a much wider range of situations.



